🧪 SIM-200 Data Reduction Guide (2×2 Binning)
Instrument: SALT RSS
Mode: SMI-200 (Slit Mask Integral Field Unit – 200μm)
Binning: 2×2
Tools: IRAF, Custom Python scripts
📁 1. Organizing Your Files
This page provides a guide to reducing Slit Mask Integral Field Unit – 200 μm (SMI-200) data with 2×2 binning using the RSS data and standard IRAF procedures. The instructions assume you are working with data processed by the SALT pipeline (referred to as “product” data).
The SALT primary reduced product data sets are generated by the in-house PySALT pipeline. This pipeline mosaics individual CCD exposures into a single FITS file, corrects for crosstalk effects, and applies bias and gain corrections.
First, it’s always important to review the Astronomer’s Log, which can be found in the /doc folder. Below are the relevant entries for the data we will be reducing:
* 01:21 Point Block ID: XXXXXX Priority: XXXXXX Proposal: XXXXXXXXXXXXXXXXXXXX Target: XXXXXXXXXXXX Instr: XXXXXXXXXXXX PI: XXXXXXXXXXXX * 01:22 Transparency: Clear External seeing: ~1.3-1.6" Guider FWHM: ~1.4" S14: Acquisition S15: Slitview P59-60: 2x1050.0s Science PG2300 SMI-200 P61-63: 3x(8.96x5 = 44.8)s Arc Ar P64-68: 5x7.0s Flat Field QTH1 and QTH2 (ND=0) * 01:26 Track Start * 02:19 Track End
You can see that they took an image of the field (S14), then an image of the slit (S15), followed by two spectra (P59-P60) with the PG2300 grating and SMI-200. For calibration, they took three arc (P61-P63) exposures of 44.8 seconds each using the Argon arc lamp, and five (P64-P68) flat-field images with QTH1 and QTH2 lamps.
And then, let’s see if this is true in the Observation Sequence:
dfits mbxgp* | fitsort object grating lampid exptime CAMANG GR-ANGLE MASKID CCDSUM CALND FILE OBJECT GRATING LAMPID EXPTIME CAMANG GR-ANGLE MASKID CCDSUM CALND mbxgpP202501070059.fits NGC3110 PG2300 NONE 1050.143 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070060.fits NGC3110 PG2300 NONE 1050.146 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070061.fits ARC PG2300 Ar 44.93 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070062.fits ARC PG2300 Ar 44.929 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070063.fits ARC PG2300 Ar 44.93 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070064.fits FLAT PG2300 QTH1 - QTH2 7.129 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070065.fits FLAT PG2300 QTH1 - QTH2 7.13 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070066.fits FLAT PG2300 QTH1 - QTH2 7.129 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070067.fits FLAT PG2300 QTH1 - QTH2 7.129 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpP202501070068.fits FLAT PG2300 QTH1 - QTH2 7.13 94.00 47.000000 PF0200N001 2 2 0.00 mbxgpS202501070014.fits NGC3110 NONE 1.07699990272522 4 4 0.00 mbxgpS202501070015.fits NGC3110 NONE 1.06900000572205 4 4 0.00
Since the images are in multi-extension format, our first step is to use imcopy to convert the images into a format that makes IRAF reduction easier.
So, in IRAF, we can run the imcopy task as shown below:
ecl> imcopy mbxgpP202501070059.fits[1][*,*][inherit+] imcopy_mbxgpP202501070059.fits ecl> imcopy mbxgpP202501070060.fits[1][*,*][inherit+] imcopy_mbxgpP202501070060.fits ecl> imcopy mbxgpP202501070061.fits[1][*,*][inherit+] imcopy_mbxgpP202501070061.fits ecl> imcopy mbxgpP202501070062.fits[1][*,*][inherit+] imcopy_mbxgpP202501070062.fits ecl> imcopy mbxgpP202501070063.fits[1][*,*][inherit+] imcopy_mbxgpP202501070063.fits ecl> imcopy mbxgpP202501070064.fits[1][*,*][inherit+] imcopy_mbxgpP202501070064.fits ecl> imcopy mbxgpP202501070065.fits[1][*,*][inherit+] imcopy_mbxgpP202501070065.fits ecl> imcopy mbxgpP202501070066.fits[1][*,*][inherit+] imcopy_mbxgpP202501070066.fits ecl> imcopy mbxgpP202501070067.fits[1][*,*][inherit+] imcopy_mbxgpP202501070067.fits ecl> imcopy mbxgpP202501070068.fits[1][*,*][inherit+] imcopy_mbxgpP202501070068.fits
It is important to know that we will trace the fibers’ positions. Fiber extraction is performed using the flat field frames. Before performing the extraction, we need to combine all the taken flats (5) into one master flat field frame. To do this, we use the imcombine task, as shown below:
Let’s start by creating these lists:
ecl>ls imcopy_mbxgpP202501070064.fits imcopy_mbxgpP202501070065.fits imcopy_mbxgpP202501070066.fits imcopy_mbxgpP202501070067.fits imcopy_mbxgpP202501070068.fits > flat.lst
Let’s now run imcombine with the following settings:
ecl> imcombine @flat.lst Master_flat.fits combine=median reject=avsigclip Jul 14 13:45: IMCOMBINE combine = median, scale = none, zero = none, weight = none reject = avsigclip, mclip = yes, nkeep = 1 lsigma = 3., hsigma = 3. blank = 0. Images imcopy_mbxgpP202501070064.fits imcopy_mbxgpP202501070065.fits imcopy_mbxgpP202501070066.fits imcopy_mbxgpP202501070067.fits imcopy_mbxgpP202501070068.fits Output image = Master_flat.fits, ncombine = 5
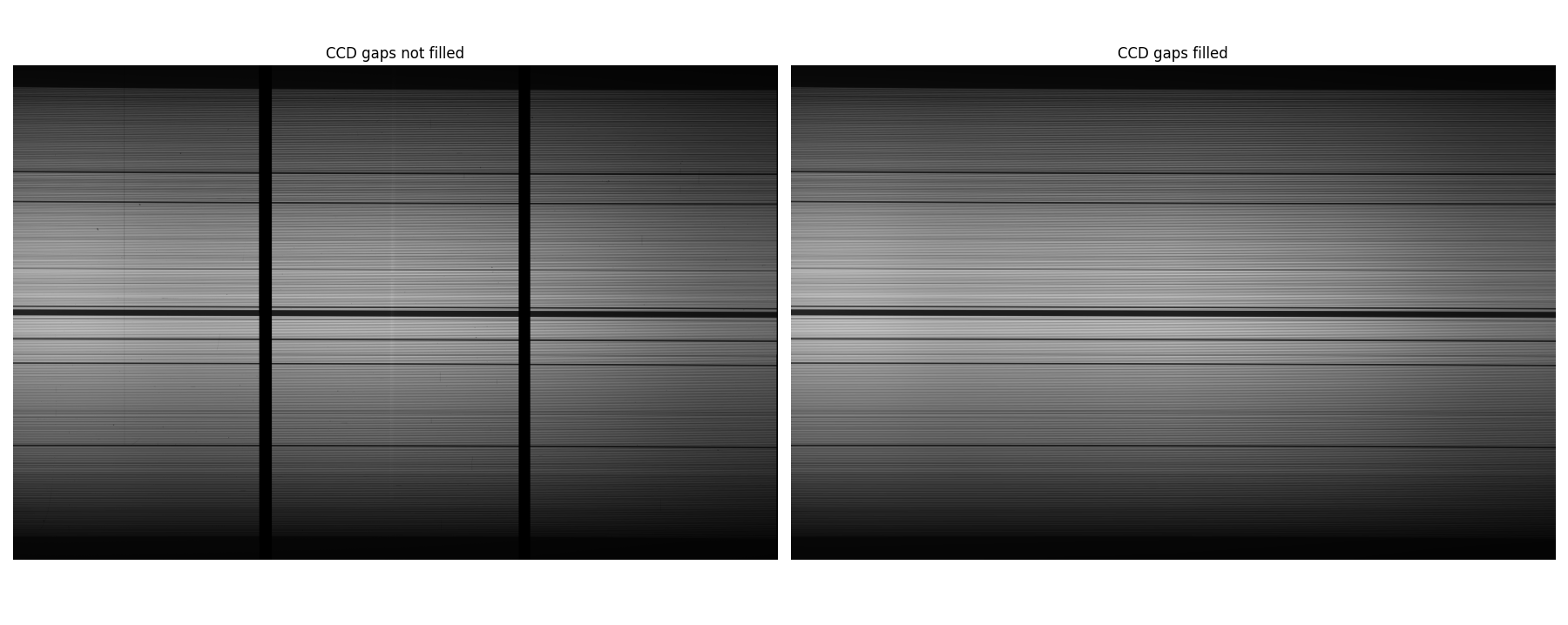
RSS has two chip gaps. To make fiber tracing efficient, we need to fit the flat field to interpolate across the chip gaps using fit1d in the image.imfit task:
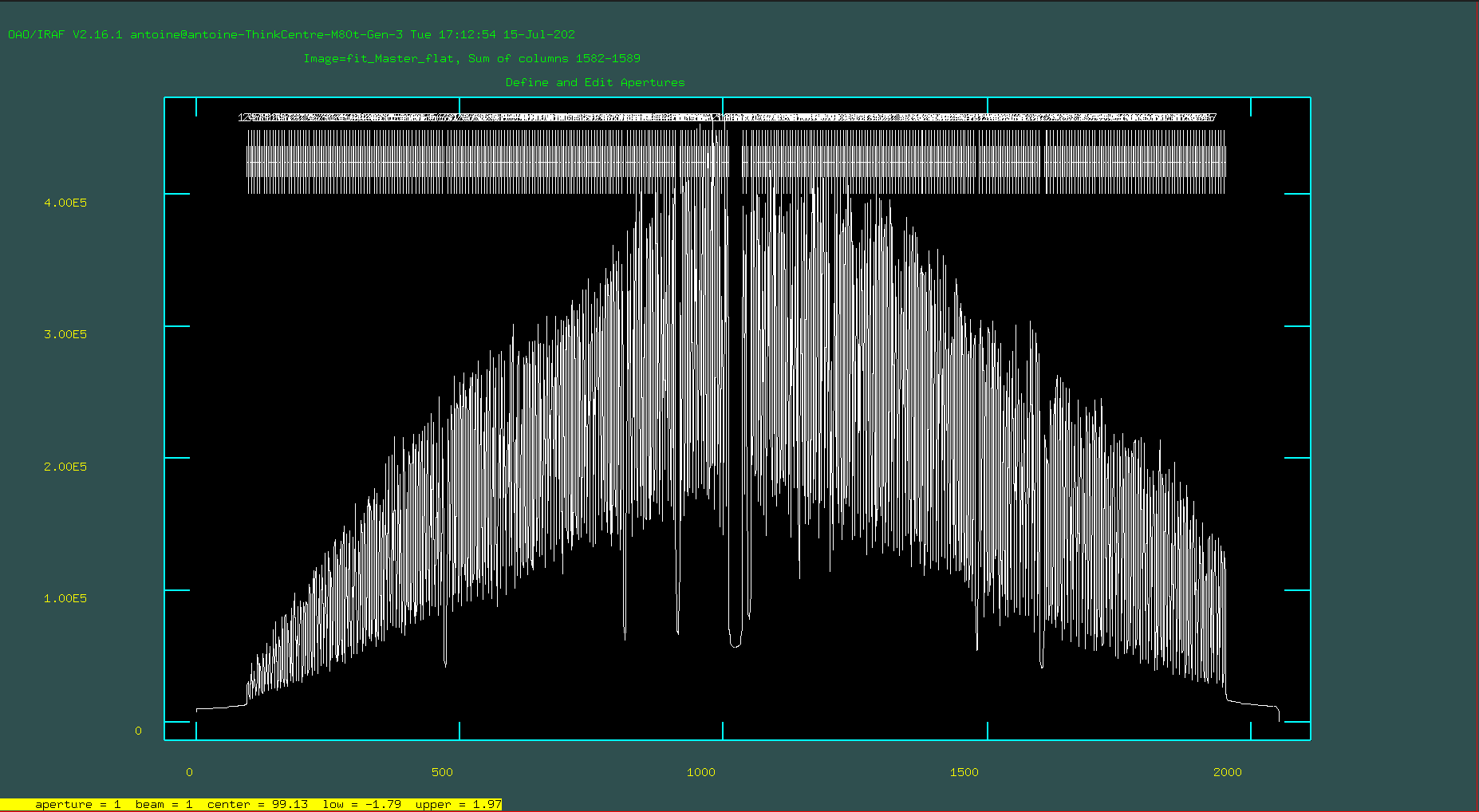
imfit> fit1d Master_flat.fits fit_Master_flat.fits fit axis=1 inter+ func=spline3 order=24 low=3 hi=3 nit=2 grow=0
Close the graphical interface window by pressing the “return” key. Take a look at the resulting image. It should look something like this:

🧼 2. Cosmic ray removal
We use the lacosmic IRAF task. You can find more information on the website L.A.Cosmic . Before you use lacosmic, you will have to load the stsdas package. Then you can look at the parameters of lacosmic using epar.
stsdas> epar lacos_spec I R A F Image Reduction and Analysis Facility PACKAGE = clpackage TASK = lacos_spec input = imcopy_mbxgpP202501070059.fits input spectrum output = imcopy_mbxgpP202501070059_cr.fits cosmic ray cleaned output spectrum outmask = Mask_imcopy_mbxgpP202501070059.fits output bad pixel map (.pl) (gain = 1.) gain (electrons/ADU) (readn = 1.) read noise (electrons) (xorder = 3) order of object fit (0=no fit) (yorder = 3) order of sky line fit (0=no fit) (sigclip = 4.5) detection limit for cosmic rays (sigma) (sigfrac = 0.5) fractional detection limit for neighbouring pixels (objlim = 1.) contrast limit between CR and underlying object (niter = 2) maximum number of iterations (verbose = yes) (mode = al)
Experiment with one of the longer exposure object frames first to find a set of parameters that does a decent job. You may want to increase niter, order of object fit or order of sky line fit . It is best to use ds9 to blink the input and output frames together.
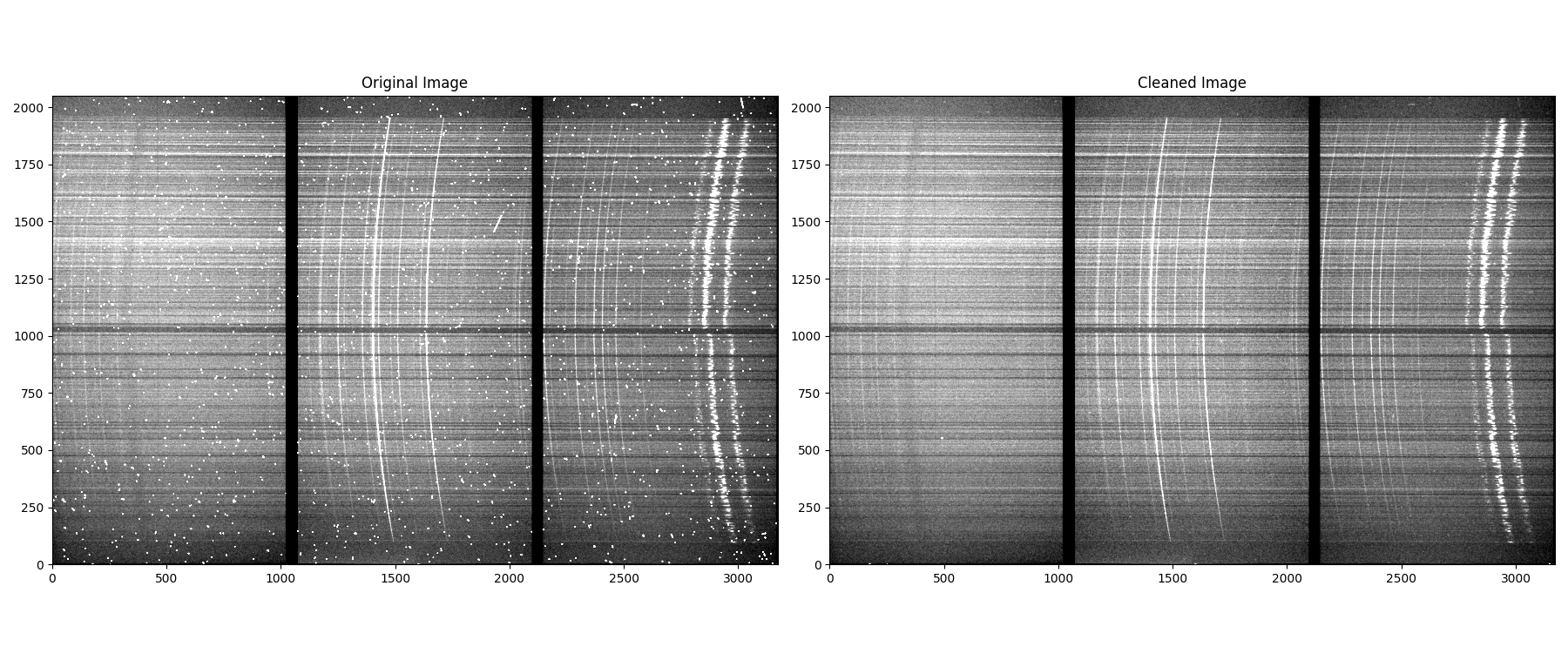
Be careful to check that lacosmic does not remove large chunks or emission lines of your spectrum. If this occurs, try increasing the value of the sigclip parameter. Reduce sigfrac and objlim to get more thorough clean. Below we show an example image:
🧵 3. Fiber Tracing & Wavelength Calibration
3.1 Trace Fibers
Aperture or fiber tracing is done with hydra.dohydra task. Be aware that SMI-200 has a total of 327 fibers (apertures) that are operational. You can set any number of “Number of fibers” as you wish, but it should be between 327 and 334. It is better to set a number above the number of working fibers (i.e., > 327).
For aperture extraction, you need to have the SMI200.iraf file.
To extract the apertures, run the dohydra task in the hydra package. Refer to the hydra.dohydra task and hydra.params task parameter files below. The important parameters to set are shown in red.
Below are the values for the ‘params’ file. Please note that these values change depending on the instrument configuration used. After modifying the parameters, type :wq to save the new settings and quit.
hydra> epar params I R A F Image Reduction and Analysis Facility PACKAGE = hydra TASK = params (line = INDEF) Default dispersion line (nsum = 8) Number of dispersion lines to sum or median (order = increasing) Order of apertures (extras = no) Extract sky, sigma, etc.? -- DEFAULT APERTURE LIMITS -- (lower =-4.) Lower aperture limit relative to center (upper =4.) Upper aperture limit relative to center -- AUTOMATIC APERTURE RESIZING PARAMETERS -- (ylevel =0.3) Fraction of peak or intensity for resizing -- TRACE PARAMETERS -- (t_step =8) Tracing step (t_funct = spline3) Trace fitting function (t_order =8) Trace fitting function order (t_niter = 1) Trace rejection iterations (t_low = 3.) Trace lower rejection sigma (t_high = 3.) Trace upper rejection sigma -- SCATTERED LIGHT PARAMETERS -- (buffer = 0.) Buffer distance from apertures (apscat1 = ) Fitting parameters across the dispersion (apscat2 = ) Fitting parameters along the dispersion -- APERTURE EXTRACTION PARAMETERS -- (weights = none) Extraction weights (none|variance) (pfit = fit1d) Profile fitting algorithm (fit1d|fit2d) (lsigma = 3.) Lower rejection threshold (usigma = 3.) Upper rejection threshold (nsubaps = 1) Number of subapertures -- FLAT FIELD FUNCTION FITTING PARAMETERS -- (f_inter = yes) Fit flat field interactively? (f_funct = spline3) Fitting function (f_order = 5) Fitting function order -- ARC DISPERSION FUNCTION PARAMETERS -- (thresho = 10.) Minimum line contrast threshold (coordli = Xe.txt) Line list (match = -3.) Line list matching limit in Angstroms (fwidth = 6.) Arc line widths in pixels (cradius = 10.) Centering radius in pixels (i_funct = spline3) Coordinate function (i_order = 1) Order of dispersion function (i_niter = 2) Rejection iterations (i_low = 3.) Lower rejection sigma (i_high = 3.) Upper rejection sigma (refit = no) Refit coordinate function when reidentifying? (addfeat = no) Add features when reidentifying? -- AUTOMATIC ARC ASSIGNMENT PARAMETERS -- (select = interp) Selection method for reference spectra (sort = ) Sort key (group = ) Group key (time = no) Is sort key a time? (timewra = 17.) Time wrap point for time sorting -- DISPERSION CORRECTION PARAMETERS -- (lineari = yes) Linearize (interpolate) spectra? (log = no) Logarithmic wavelength scale? (flux = yes) Conserve flux? -- SKY SUBTRACTION PARAMETERS -- (combine = average) Type of combine operation (reject = avsigclip) Sky rejection option (scale = none) Sky scaling option (mode = ql)
Here are the values for the ‘dohydra’ file. Note that these parameters need to be changed for different instrument configurations. The apidtab file can be found here. Download and save it to your working directory.
hydra> epar dohydra I R A F Image Reduction and Analysis Facility PACKAGE = hydra TASK = dohydra objects =imcopy_mbxgpP202501070061_cg_cr.fitsList of object spectra (apref =fit_Master_flat.fits) Aperture reference spectrum (flat = ) Flat field spectrum (through = ) Throughput file or image (optional) (arcs1 = ) List of arc spectra (arcs2 = ) List of shift arc spectra (arcrepl = ) Special aperture replacements (arctabl = ) Arc assignment table (optional) (readnoi = 2.3) Read out noise sigma (photons) (gain = 1.5) Photon gain (photons/data number) (datamax = INDEF) Max data value / cosmic ray threshold (fibers =328) Number of fibers (width =4.) Width of profiles (pixels) (minsep =4.) Minimum separation between fibers (pixels) (maxsep =20.) Maximum separation between fibers (pixels) (apidtab =SMI200.iraf) Aperture identifications (crval = INDEF) Approximate central wavelength (cdelt = INDEF) Approximate dispersion (objaps = ) Object apertures (skyaps = ) Sky apertures (arcaps = ) Arc apertures (objbeam = 0,1) Object beam numbers (skybeam = 0) Sky beam numbers (arcbeam = ) Arc beam numbers (scatter = no) Subtract scattered light? (fitflat = no) Fit and ratio flat field spectrum? (clean = no) Detect and replace bad pixels? (dispcor = no) Dispersion correct spectra? (savearc = no) Save simultaneous arc apertures? (skyalig = no) Align sky lines? (skysubt = no) Subtract sky? (skyedit = no) Edit the sky spectra? (savesky = no) Save sky spectra? (splot = no) Plot the final spectrum? (redo = no) Redo operations if previously done? (update = no) Update spectra if cal data changes? (batch = no) Extract objects in batch? (listonl = no) List steps but don't process? (params = ) Algorithm parameters (mode = ql)
If you are happy with all changes made to the parameters, type :wq to save the new settings and quit.
While running dohydra, on the first run, if it asks you’d like to so something, always say yes (see below):
hydra> dohydra List of object spectra (imcopy_mbxgpP202501070061_cg_cr.fits): Set reference apertures for fit_Master_flat Resize apertures for fit_Master_flat? (yes): Edit apertures for fit_Master_flat? (yes):
You can zoom in to see the aperture identification by pressing the w and e keys, then pressing e again to select the region you wish to zoom into. To zoom out, press w followed bya. See below for an example of how it looks:
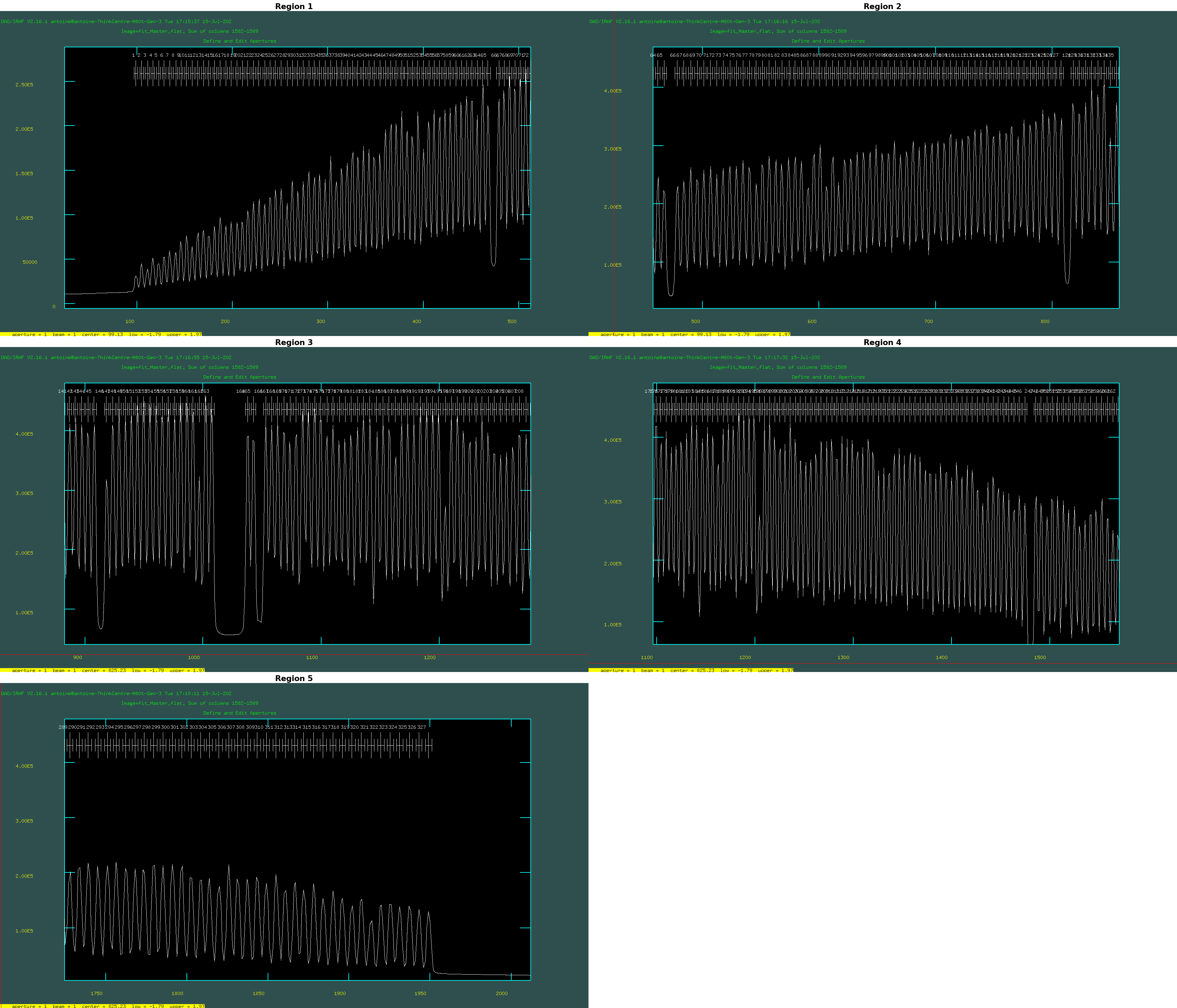
If you are happy with all traced apertures press the "q" key. Fit traced positions for flat_gap_int_gal interactively? (yes): no
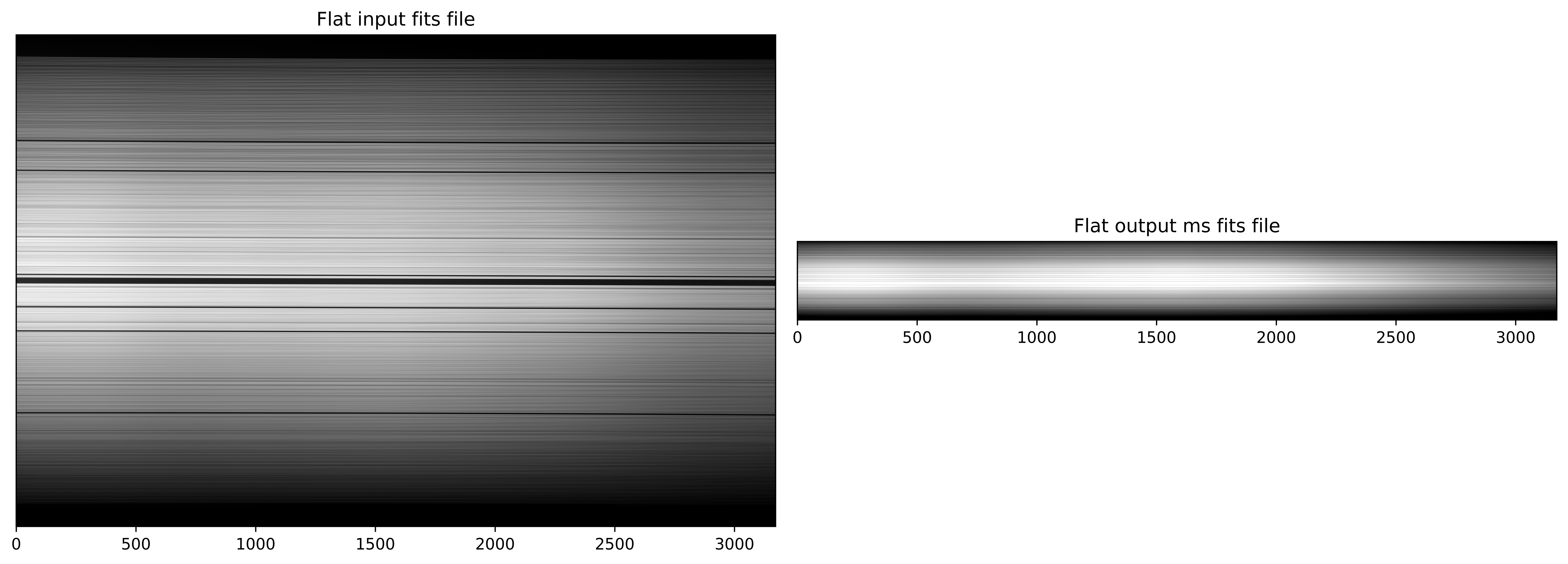
Next, dohydra will fit (trace) the apref images (flat). Use an order 8 spline3 function, and force dohydra to use the same fitting fuction for all fibres.
Take a look at the result. If it looks all right, we’ll move on.


So, now we’re going to do the only difficult portion of this reduction procedure: determining the wavelength solution—the part where we identify emission lines in the arc lamp spectrum for the purposes of wavelength calibration. The way in which this works is that we use an IRAF task called identify, which first plots the centers of the arc image, showing the peaks, and then allows the user to provide the wavelengths of those peaks. From here, a fit will be performed between the pixel values and the wavelength values for the peaks near the center of the image, and you will move on to the tasks reidentify and dispcor. The task used for wavelength calibration is in the onedspec package.
3.2 Arc Line Identification
There are a few important things to notice from how we ran identify:
First, we need to make sure that the coordlist is set to the file you should have downloaded, corresponding to that particular arc lamp. The SALT arc lamp files can be downloaded here.
We start with the identify task, and below are the parameters to set (in red).
onedspec> epar identify I R A F Image Reduction and Analysis Facility PACKAGE = onedspec TASK = identify images =imcopy_mbxgpP202501070061_cg_cr.fitsImages containing features to be identified (section = middle line) Section to apply to two dimensional images (databas = database) Database in which to record feature data (coordli =path to the arc lamp text file Ar_PG2300.txt) User coordinate list (units = ) Coordinate units (nsum =1) Number of lines/columns/bands to sum in 2D image (match = -3.) Coordinate list matching limit (maxfeat =50) Maximum number of features for automatic identif (zwidth = 100.) Zoom graph width in user units (ftype = emission) Feature type (fwidth =6.) Feature width in pixels (cradius = 5.) Centering radius in pixels (thresho =0.) Feature threshold for centering (minsep = 2.) Minimum pixel separation (functio =spline3) Coordinate function (order =1) Order of coordinate function (sample = *) Coordinate sample regions (niterat = 0) Rejection iterations (low_rej = 3.) Lower rejection sigma (high_re = 3.) Upper rejection sigma (grow = 0.) Rejection growing radius (autowri = no) Automatically write to database (graphic = stdgraph) Graphics output device (cursor = ) Graphics cursor input crval = Approximate coordinate (at reference pixel) cdelt = Approximate dispersion (aidpars = ) Automatic identification algorithm parameters (mode = ql)
Now, when you run that identify line, you’ll be presented with the spectrum of the emission features from the center of the image (the center row, which corresponds to row 164).
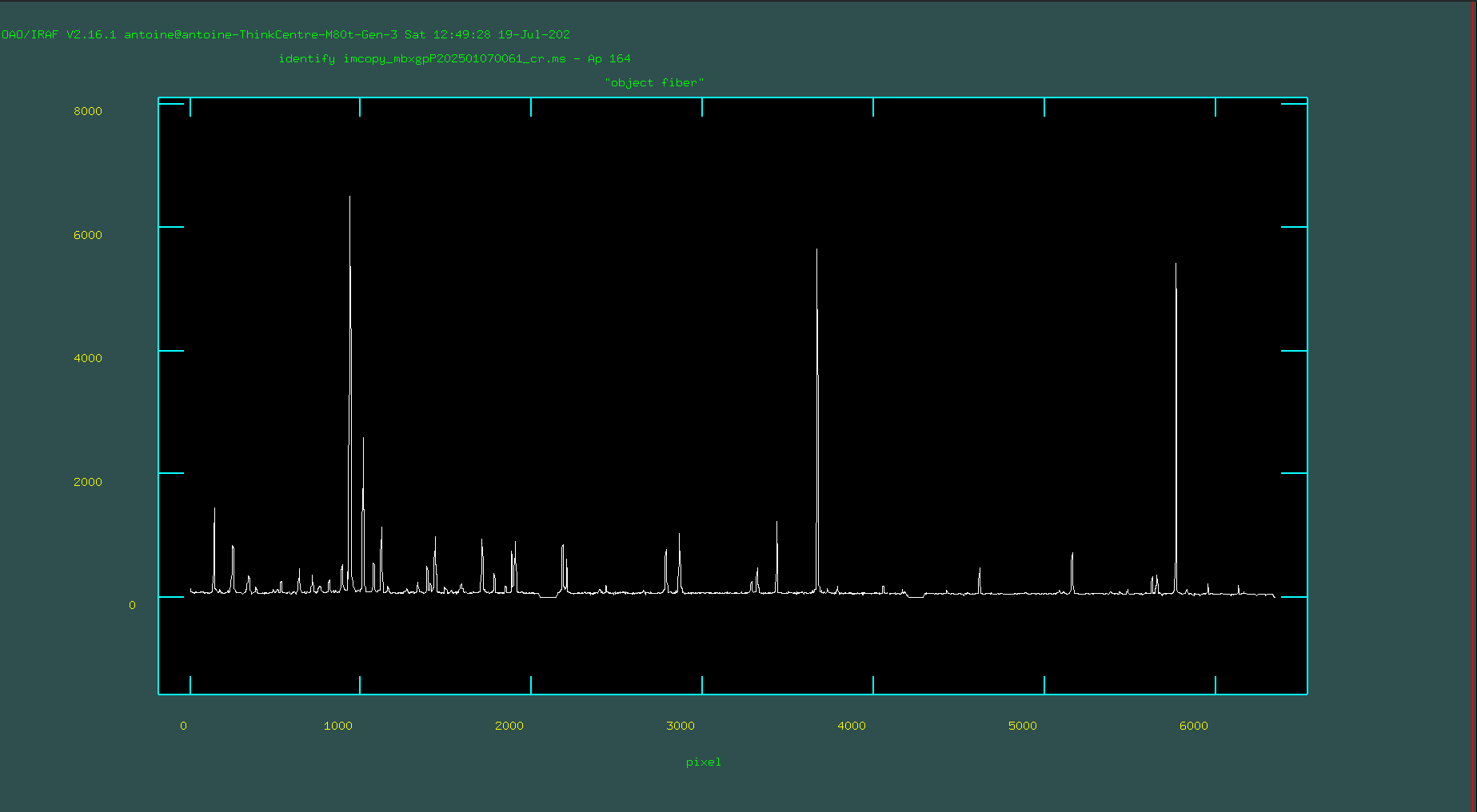
Your task is to go through and identify the lines you see in your sample arc spectrum and mark them. To do this, you’ll want to zoom in on a region of interest in the arc. To zoom in, make sure you’ve clicked on the pop-up window that shows the arc spectrum and type ‘w’ (for “window”). The main objective here is to identify 4–6 lines spanning the pixel range, with values found in the reference library. These form the initial solution, from which additional lines are identified, leading to a final, refined solution.
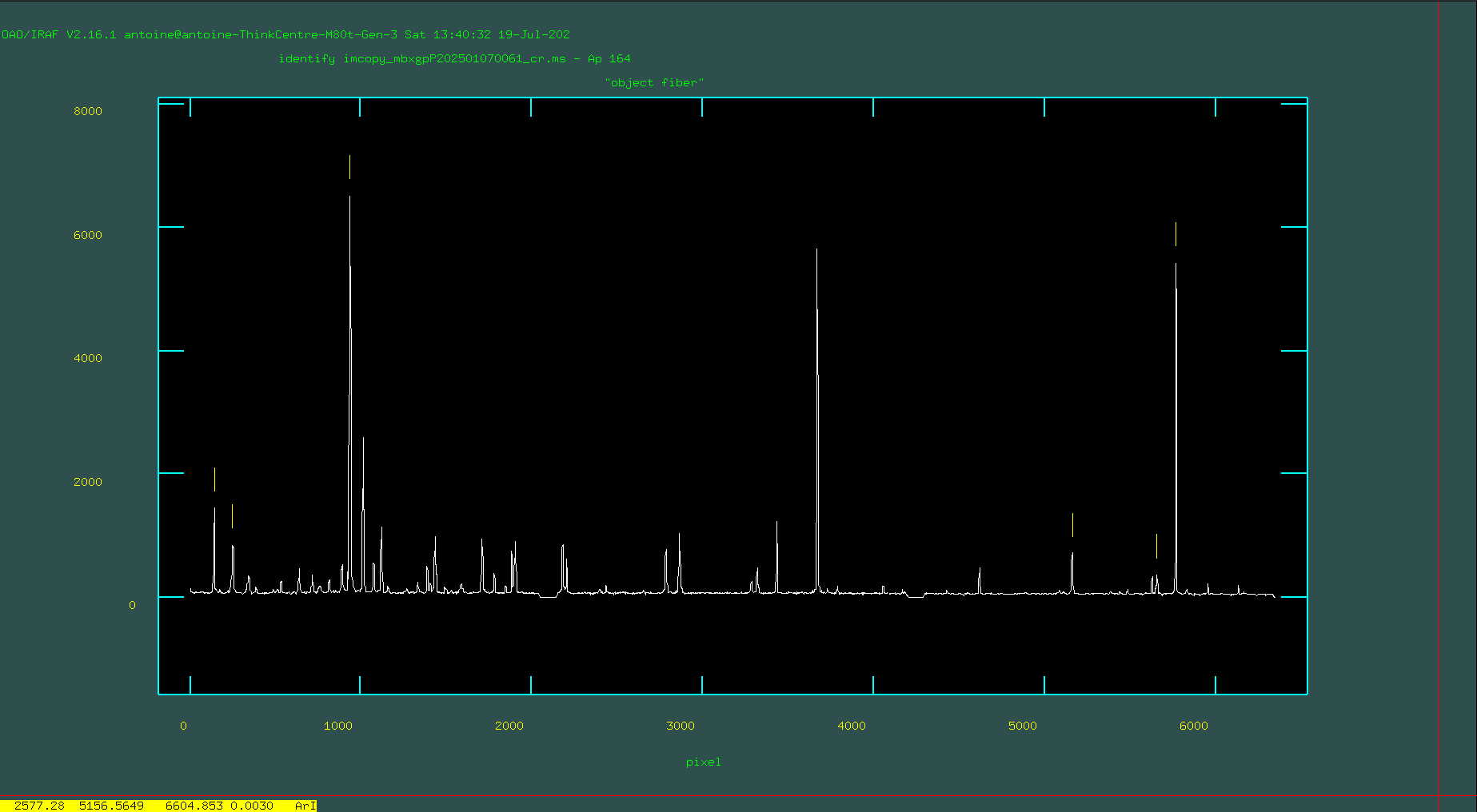
Next, type “f” to fit these 4–6 points. If possible, fit with an order 1 spline3 function. Only increase the order if absolutely necessary. If the lines are marked correctly, you should have a very low RMS value. If not, hit “q” to quit the fit, erase all the lines you marked, and then redo the markings and refit with “f”.
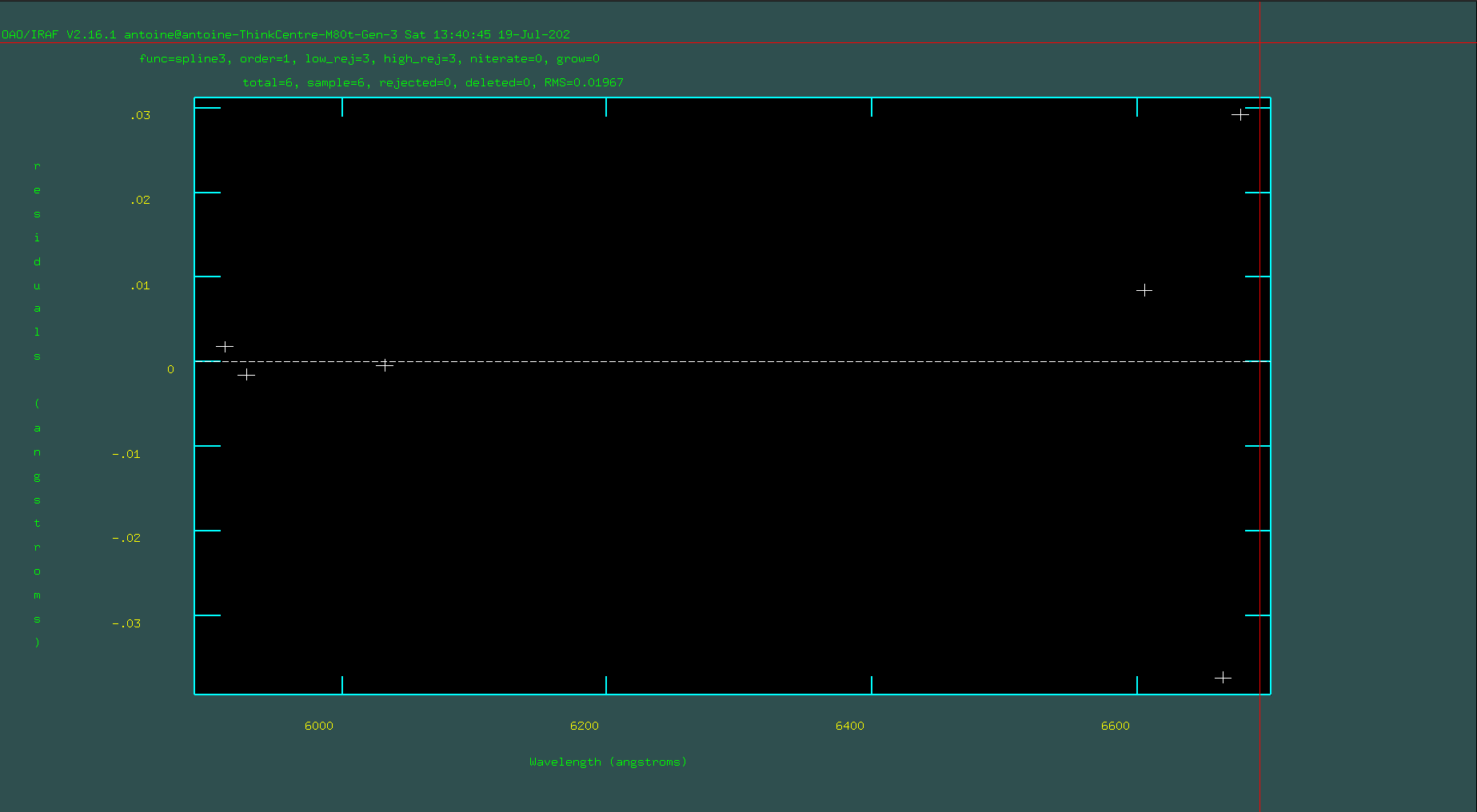
Once you’ve achieved a good initial fit, return to the spectrum-plotting and marking screen by pressing “q,” and then type “l” to read other lines from the line list and mark lines near the peaks in the spectrum. This step is crucial because it provides a high-precision list of lines.
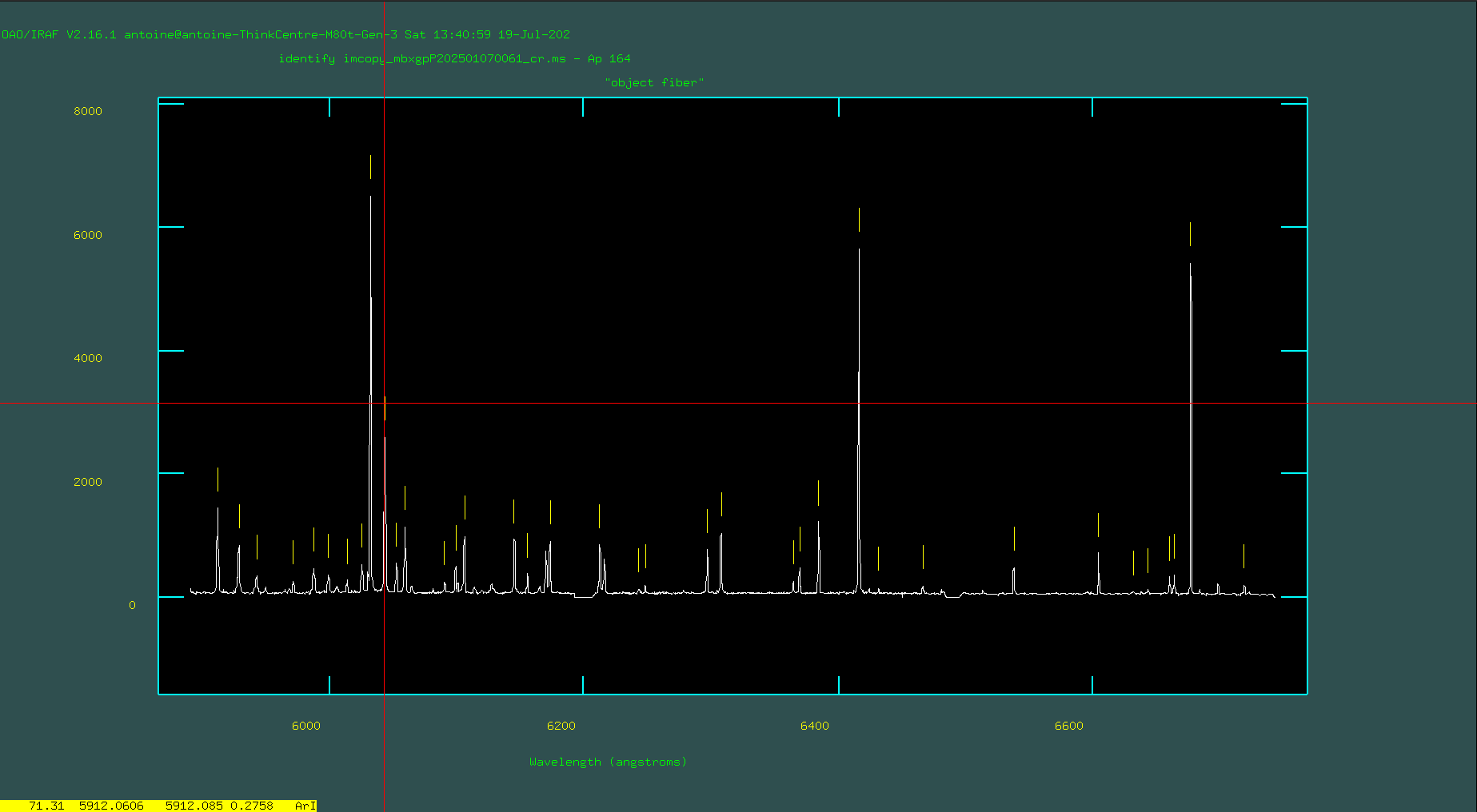
Hit “f” to fit again. This time, when fitting, check to see which lines are outliers by flipping between “f” and “q.” Delete outliers with “d” either in the spectrum plot or the fitted residuals plot, and delete any “lines” that have been automatically marked and appear to have low S/N or might be on blended lines.
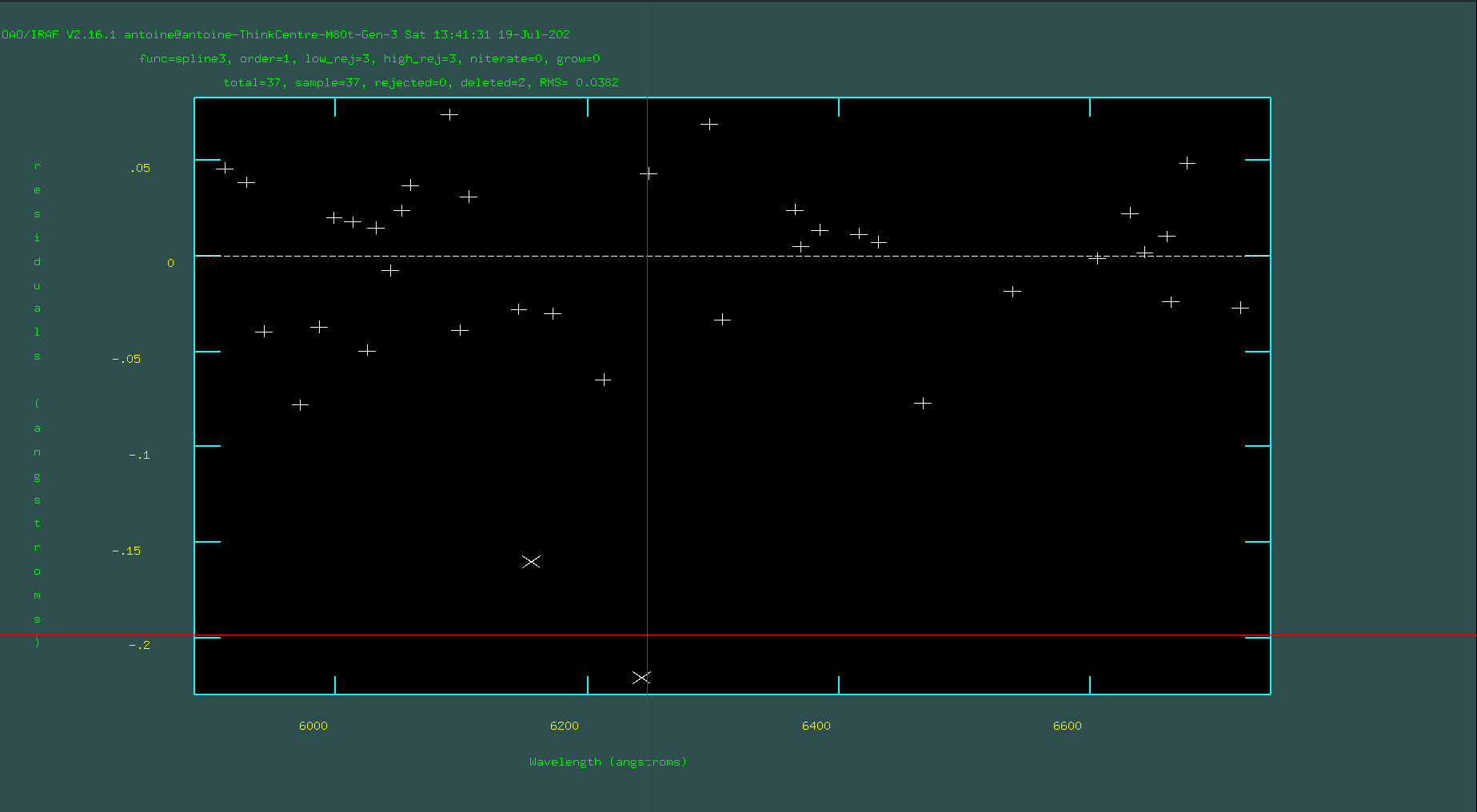
Iterate until you’re statisfied with the fit: Pay attention to RMS, and when the fit is good and RMS is small enough (rms <=0.05).
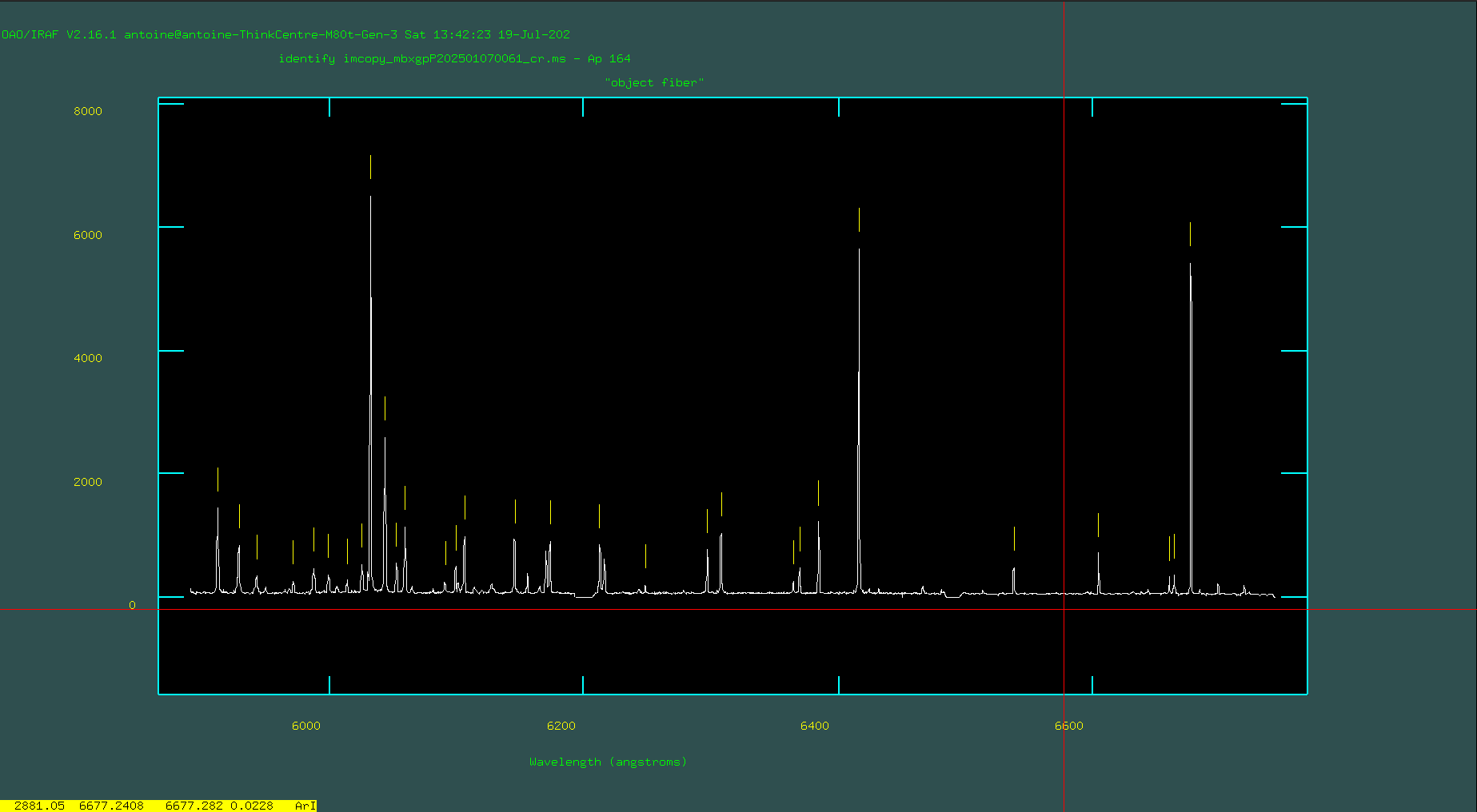
After fitting the center row, quit the graphical interface by pressing ‘q’ and will ask you if you want to “Write the feature data to the database (yes)?” Here you should answer “yes”.
After running identify successfully now you have to go to the next step which is to run a reidentify task. Below are the parameters to set (in red):
onedspec> epar reidentify I R A F Image Reduction and Analysis Facility PACKAGE = onedspec TASK = reidentify referenc =imcopy_mbxgpP202501070061_cg_cr.fitsReference image images =imcopy_mbxgpP202501070061_cg_cr.fitsImages to be reidentified (interac = no) Interactive fitting? (section =middle line)Section to apply to two dimensional images (newaps = yes) Reidentify apertures in images not in reference? (overrid =yes) Override previous solutions? (refit =yes) Refit coordinate function? (trace =yes) Trace reference image? (step =1) Step in lines/columns/bands for tracing an image (nsum =1) Number of lines/columns/bands to sum (shift =INDEF) Shift to add to reference features (INDEF to search) (search =INDEF) Search radius (nlost =35) Maximum number of features which may be lost (cradius = 5.) Centering radius (thresho = 0.) Feature threshold for centering (addfeat =no) Add features from a line list? (coordli =path to the arc lamp text file Ar_PG2300.txt) User coordinate list (match = -3.) Coordinate list matching limit (maxfeat =50) Maximum number of features for automatic identification (minsep = 2.) Minimum pixel separation (databas = database) Database (logfile = logfile) List of log files (plotfil = ) Plot file for residuals (verbose = no) Verbose output? (graphic = stdgraph) Graphics output device (cursor = ) Graphics cursor input answer = yes Fit dispersion function interactively? crval = Approximate coordinate (at reference pixel) cdelt = Approximate dispersion (aidpars = ) Automatic identification algorithm parameters (mode = ql)
It may appear that nothing is happening visually, but the id* files in the database/ directory will have been updated accordingly. These files contain identification data that the system uses for subsequent processing.
Important: Monitor the progress of the re-identification task to ensure it successfully identifies all 327 apertures in total. You can verify this by checking the output logs or the status within the system’s interface.
If, after the task completes, not all apertures are identified (i.e., fewer than 327 are recognized), you’ll need to run the identification task again. This ensures that the system correctly associates all apertures with their respective identities, which is crucial for accurate data analysis and results.
The final step is to run a “dispcor” task to apply a dispersion solution, rectify and resample data. Below are the parameters to set (in red):
onedspec> epar dispcor I R A F Image Reduction and Analysis Facility PACKAGE = onedspec TASK = dispcor input =imcopy_mbxgpP202501070061_cr.ms.fitsList of input spectra output =imcopy_mbxgpP202501070061_cr.msw.fitsList of output spectra (lineari =yes) Linearize (interpolate) spectra? (databas = database) Dispersion solution database (table = ) Wavelength table for apertures (w1 = INDEF) Starting wavelength (w2 = INDEF) Ending wavelength (dw = INDEF) Wavelength interval per pixel (nw = INDEF) Number of output pixels (log = no) Logarithmic wavelength scale? (flux =yes) Conserve total flux? (blank = 0.) Output value of points not in input (samedis =yes) Same dispersion in all apertures? (global = no) Apply global defaults? (ignorea = no) Ignore apertures? (confirm = no) Confirm dispersion coordinates? (listonl = no) List the dispersion coordinates only? (verbose = yes) Print linear dispersion assignments? (logfile = ) Log file (mode = ql)
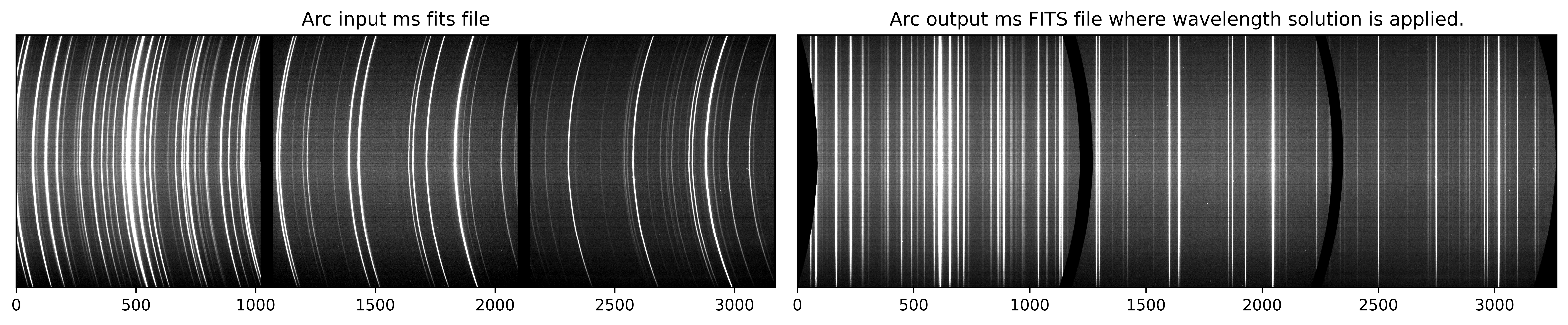
🌈 4. Apply Wavelength Solution
To apply the wavelength correction to the science file, you need to add information about the reference solution in the header, and then use ‘dispcor’ in the ‘onedspec’ package.
hedit imcopy_mbxgpP202501070059_cr.ms.fits REFSPEC1 "imcopy_mbxgpP202501070061_cr.ms" add+ ver- show+ hedit imcopy_mbxgpP202501070060_cr.ms.fits REFSPEC1 "imcopy_mbxgpP202501070061_cr.ms" add+ ver- show+
Run the dispcor task for the science file, and you are done with wavelength calibration.
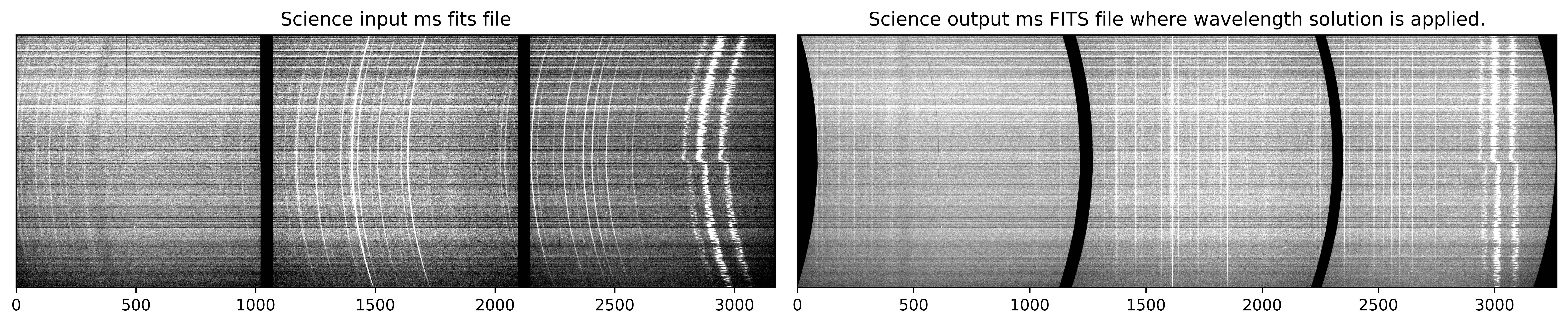
🔬 7. Flux Calibration
At this stage, you can now go ahead and perform flux calibration. Remember that with SALT, you can do relative flux calibration.
To do flux calibration you have to follow the below step:
You have to correct the instrument response by multiplying an efficiency factor to the SMI measured flux as true standard star flux. The efficiency factor consists of telescope throughput, telescope illumination, spectral response, pixel to pixel variation, slit efficiency and spectrograph vignetting. Check throughput value from SALT call for proposal document and telescope illumination provided at fits header (PUPSTA and PUPEND). The spectral response including lamp response and spectrograph response as well as pixel-to-pixel variation is taken out by dividing the object frame with the mean normalized flat (wavelength calibrated file). The slit efficiency is based on the number of fibers that are illuminated by the standard star while the spectrograph vignetting is kept at 1 since the fiber bundle is sitting at the middle of the slit.
Calculate the slit efficiency using the formula:
Slit Efficiency = Nf / ((2 * n + 1) * 1.2)2
where:
- Nf is the total number of fibers illuminated by the standard star
- n is the number of fiber rings
To get n, you have to solve the equation:
n(n + 1) / 2 = (Nf – 1) / 6
Nf can be obtained by plotting observed standard stars (sum across the columns (x-axis) to collapse the spectrum along the slit (rows, y-axis)) and checking how many fibers are illuminated. Below is the image which shows you how this has to look like.
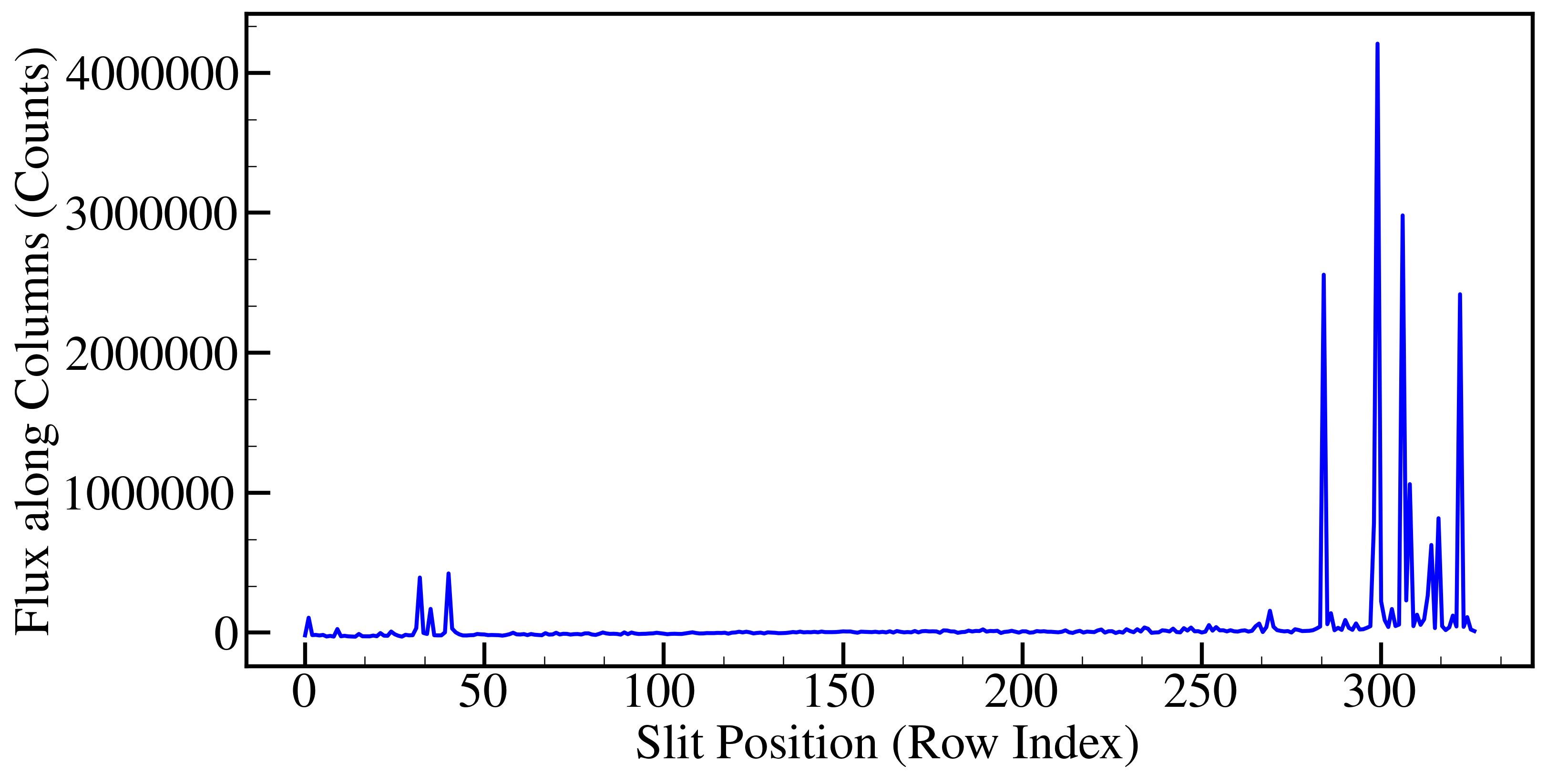
In this case, we found that N_f equals 19. Solving n(n+1)/2 = (N_f-1)/6 yields n equal to 2. After getting the values of N_f and n, you can calculate slit efficiency using equation 1.
Next is to extract all illuminated fibers from ms file and sum all into one fiber. You will end up with 1D spectral (see the image below).
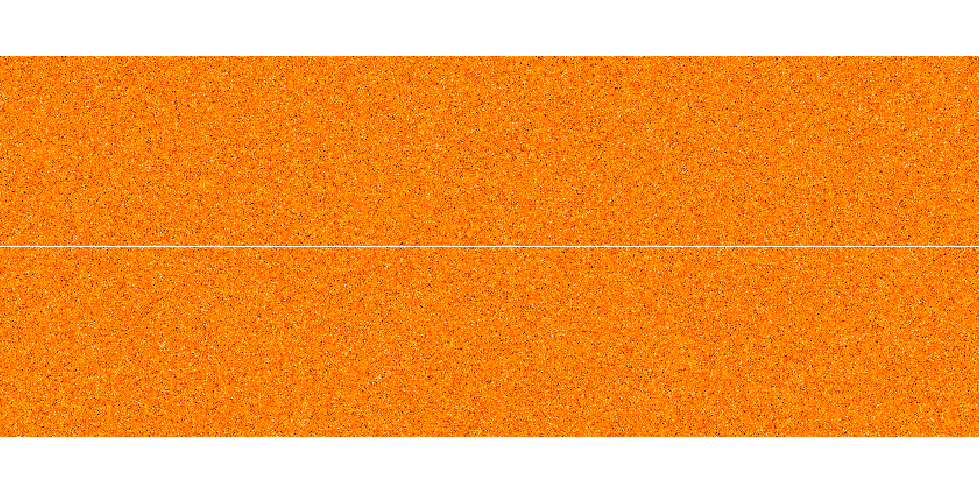

Now you have to take this 1D spectral multiple with efficiency factor (telescope throughput, telescope illumination, slit efficiency and spectrograph vignetting).
Next you need:
- A file containing the data for the observed standard star *.dat
- Check in /iraf/iraf/noao/lib/onedstds
- e.g. LTT6248 is found as l6248.dat
- However, it is not in the default spec50cal directory, but in the ctionewcal directory, so that needs to be specified in the parameter file. If star is not there, google ESO standard pages , and copy one from there, just remember to use the version in MAGNITUDES.
- What about the extinction file? You could just use suth_extinct.dat.
Check that exptime and air mass are in the fits header.
Prepare the standard star file (using noao.onedspec.standard task) and obtain the transformation function (by using noao.onedspec.sensfunc task).
I R A F Image Reduction and Analysis Facility PACKAGE = onedspec TASK = standard input =summed_rows_standr_HILT600.fitsInput image file root name output =standard_HILT600.datOutput flux file (used by SENSFUNC) (samesta= yes) Same star in all apertures? (beam_sw= no) Beam switch spectra? (apertur= ) Aperture selection list (bandwid= INDEF) Bandpass widths (bandsep= INDEF) Bandpass separation (fnuzero= 3.6800000000000E-20) Absolute flux zero point (extinct=Path to suth_extinct.dat file) Extinction file (caldir =Path to Standard stars directory) Directory containing calibration data (observa=saao) Observatory for data (interac= yes) Graphic interaction to define new bandpasses (graphic= stdgraph) Graphics output device (cursor = ) Graphics cursor input star_nam=HILT600Star name in calibration list airmass =1.266146Airmass exptime =150.13Exposure time (seconds) mag = Magnitude of star magband = Magnitude type teff = Effective temperature or spectral type answer = yes (no|yes|NO|YES|NO!|YES!) (mode = ql)
Save and run the standard task. This will open a graphical interface where you should delete points, areas, or lines outside of the continuum—in standard by pressing d. If you are done, you have to quit the graphical interface by pressing q. Outputs a file called standard_HILT600.dat in this case.
When the task is executed, the following plot will appear:
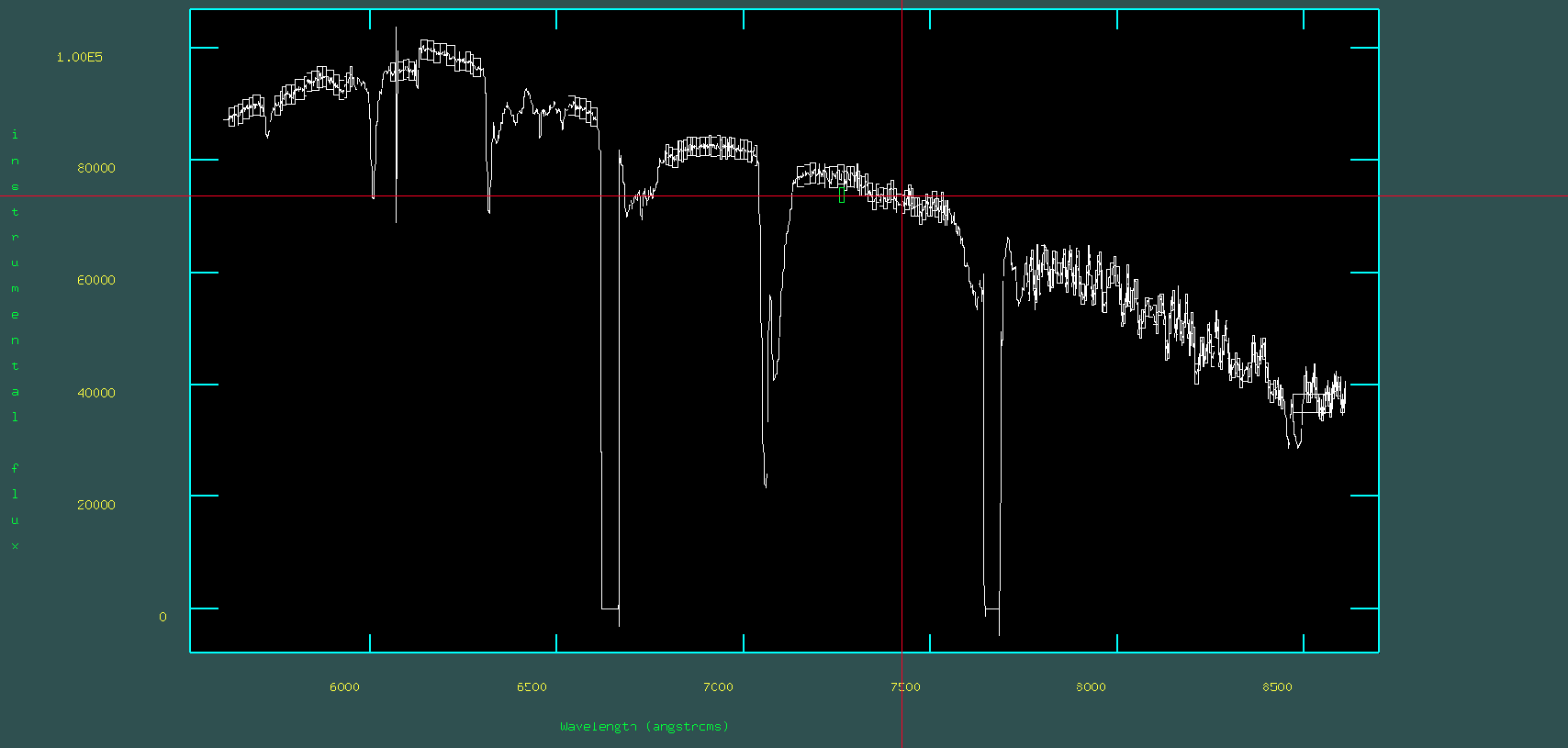
Next step is to run the sensfunc task in the onedspec package.
Fitting the sensitivity function:
Provided with all the needed information, in this step the sensitivity function will be fitted.
I R A F Image Reduction and Analysis Facility PACKAGE = onedspec TASK = sensfunc standard=standard_HILT600.datInput standard star data file (from STANDARD) sensitiv=sensfunc_HILT600.fitsOutput root sensitivity function imagename (apertur= ) Aperture selection list (ignorea= no) Ignore apertures and make one sensitivity function? (logfile= logfile) Output log for statistics information (extinct= ) Extinction file (newexti=Path to suth_extinct.dat file) Output revised extinction file (observa=saao) Observatory of data (functio= spline3) Fitting function (order = 6) Order of fit (interac= yes) Determine sensitivity function interactively? (graphs = sr) Graphs per frame (marks = plus cross box) Data mark types (marks deleted added) (colors = 2 1 3 4) Colors (lines marks deleted added) (cursor = ) Graphics cursor input (device = stdgraph) Graphics output device answer = yes (no|yes|NO|YES) (mode = ql)
When the task is executed, two plots, as shown in the figure below, will appear. The upper plot displays the data to which the sensitivity function fit is applied, along with the fit itself. As before, you can modify the fitting order and redo the fit by pressing f, delete outliers with d, and finish with q. The lower plot shows the residuals of the fit. The goal is to have the residuals normally distributed around zero along the y-axis.
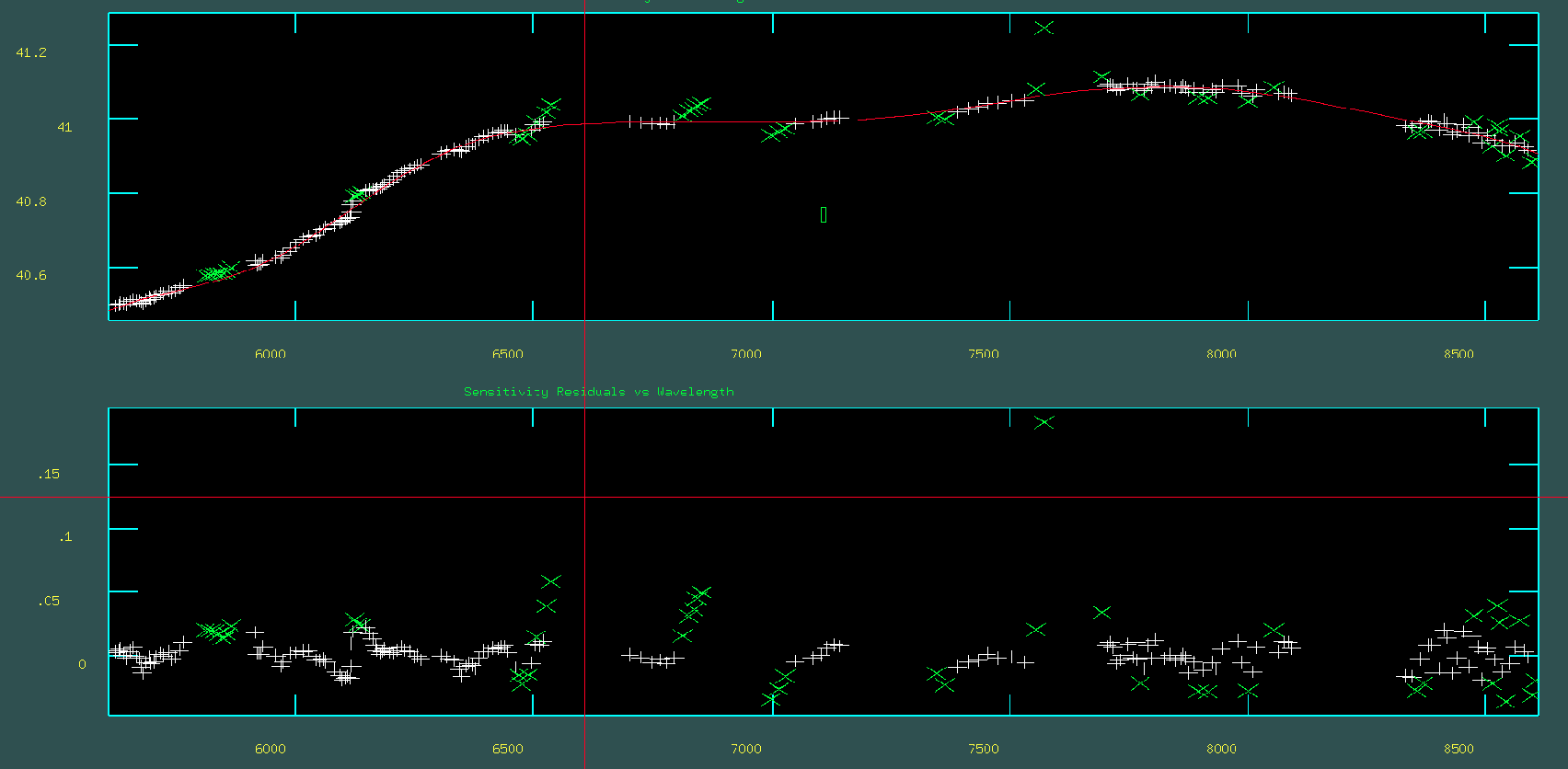
Finally flux calibration of the standard star and sciences file with noao.onedspec.calibrate function.
I R A F
Image Reduction and Analysis Facility
PACKAGE = onedspec
TASK = calibrate
input =summed_rows_standr_HILT600.fits Input spectra to calibrate
output = summed_rows_standr_HILT600_flux.fits Output calibrated spectra
(extinct= yes) Apply extinction correction?
(flux = yes) Apply flux calibration?
(extinct= Path to suth_extinct.dat file) Extinction file
(observa= saao ) Observatory of observation
(ignorea= no) Ignore aperture numbers in flux calibration?
(sensiti= Path to sensitivity spectra ) Image root name for sensitivity spectra
(fnu = no) Create spectra having units of FNU?
airmass = 1.266146 Airmass
exptime = 150.13 Exposure time (seconds)
(mode = ql)
🧊 8. Cube Reconstruction
This is a Python program that processes spectral data from SMI-200 for grid mapping and spatial reconstruction with fiber data. In the code, we use the Shepard algorithm. Developed by Donald Shepard in 1968, it is also known as inverse distance weighting interpolation. A full explanation of how the program works is given in the readme file.